How to calculate the spatial gradient of a scalar field in cartesian coordiantes#
FELiCS can be used to calculate the spatial gradient of a quantity with a few commands. Here, the following cases are described:
cartesian coordinates: square 2D domain
cartesian coordinates with spectral dimension: the de-facto 2D domain can be imagined as a 3D channel, which is periodically expanded in the third direction
Cartesian coordinates on a 2D domain#
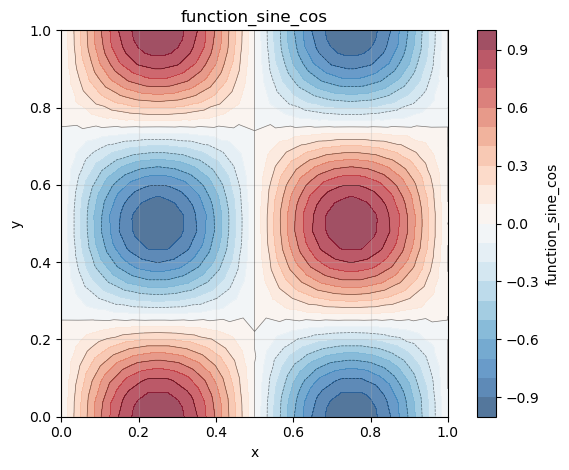
Here, the gradient of a scalar field on a square 2D mesh is computed. We define our scalar field as follows:
Step 1: Create FELiCSMesh, FELiCSSpace and Field
from FELiCS.SpaceDisc.FELiCSMesh import FELiCSMesh
from FELiCS.SpaceDisc.FEMSpaces import create_function_space
from FELiCS.Fields.Field import Field
from FELiCS.IO.Reader import Reader
meshFileName = "./square_mesh.msh"
coordinateSystemName = "Cartesian"
# create a FELiCS mesh from the gmsh file
mesh = FELiCSMesh(coordinateSystemName, meshFileName)
# create a scalar space
scalarSpace = create_function_space(mesh, degree=2, dim=1)
# create a Field object and import the data from the h5 file
phi = Field(scalarSpace, mesh=mesh, name="function_sine_cos")
phi.import_data(Reader(), "input.h5")
Info | FELiCSMesh.py | __init__ (line 124 ) : Opening mesh file: ./square_mesh.msh
Info | FELiCSMesh.py | __init__ (line 132 ) : Mesh contains 513 nodes and 1024 elements
Info | Reader.py | _interpolate_to_calc_mesh (line 706 ) : Linear interpolation took 0.6 seconds. (Mesh size: (1969, 2))
(<FELiCS.Fields.Field.Field at 0x17fee7770>, [])
Step 2: Get gradient as a vector field and extract the scalar fields
We now want to calculate the gradient of our scalar quantity using the getGradientField() method of our Field class. This method returns a Field object, which represents a vector-field as we calculate the gradient of a scalar with respect to our two spatial and one spectral dimension. The gradient is defined as
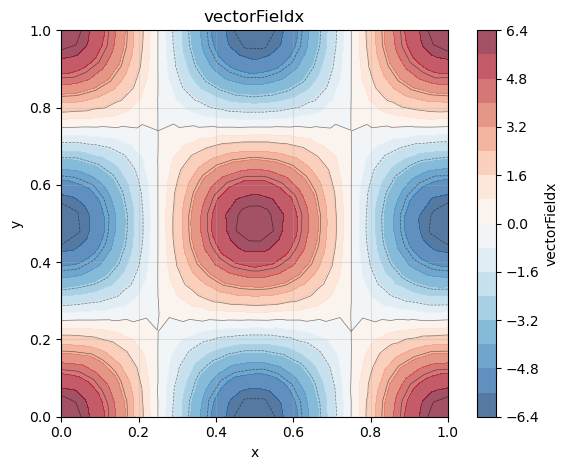
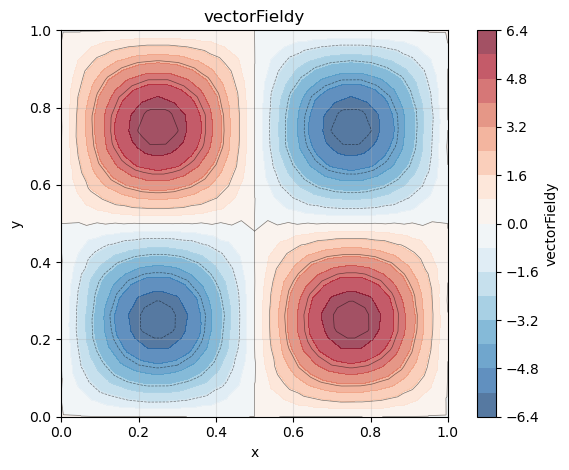
So for our function the analytical gradient is
The resulting field is defined on a vector space with 2 dimension. We can also transform it in two scalar fields via the command ‘getListOfSingleFields()’.
gradPhi = phi.get_gradient_field() # this is a vector field
# get scalar fields from gradient:
gradList = gradPhi.get_list_of_sub_fields()
phi_x = gradList[0]
phi_y = gradList[1]
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
Step 2: Postprocessing
Exporting as h5 file and visualization of the gradients by using the FELiCS plotting function:
from FELiCS.IO.Writer import Writer
gradPhi.export_to_h5(Writer())
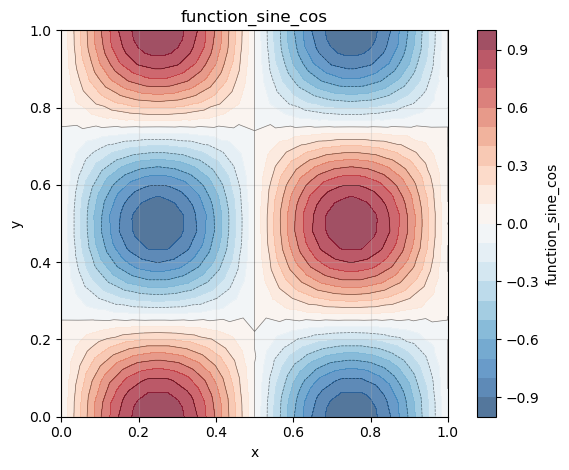
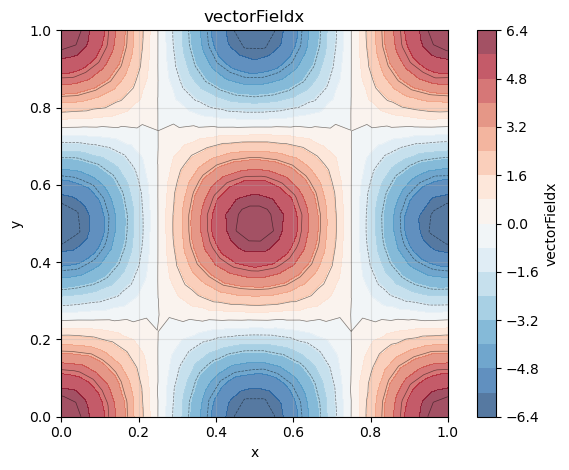
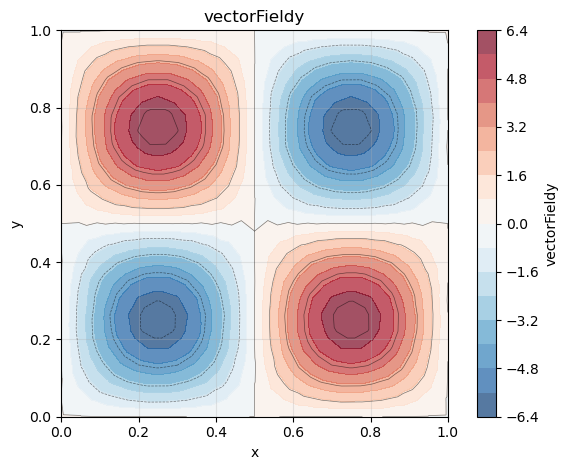
phi.plot()
phi_x.plot()
phi_y.plot()
Cartesian coordinates with spectral dimension#
Here, to the two spatial dimensions (x and y) a third spectral dimension in z-direction is added. With the spectral dimension the scalar field becomes
where m is the wave number. With the wave number we represent a periodicity of the flow in z-direction, while still only calculating a 2D flowfield.
Step 1: Create FELiCSMesh, FELiCSSpace and Field
from FELiCS.SpaceDisc.FELiCSMesh import FELiCSMesh
from FELiCS.SpaceDisc.FEMSpaces import create_function_space
from FELiCS.Fields.Field import Field
from FELiCS.IO.Reader import Reader
meshFileName = "./square_mesh.msh"
coordinateSystemName = "Cartesian"
m = 3
# create a FELiCS mesh from the gmsh file
# by setting m to an integer number that is not zero, the spectral direction is added to the coordinate system
mesh = FELiCSMesh(coordinateSystemName, meshFileName, m=m)
# create a scalar space
scalarSpace = create_function_space(mesh, degree=2, dim=1)
# create a Field object and import the data from the h5 file
# by setting m to an integer number that is not zero, the spectral direction is added to the field
phi = Field(scalarSpace, mesh=mesh, name = "function_sine_cos", m = m)
phi.import_data(Reader(),"input.h5")
Info | FELiCSMesh.py | __init__ (line 124 ) : Opening mesh file: ./square_mesh.msh
Info | FELiCSMesh.py | __init__ (line 132 ) : Mesh contains 513 nodes and 1024 elements
Info | Reader.py | _interpolate_to_calc_mesh (line 706 ) : Linear interpolation took 0.6 seconds. (Mesh size: (1969, 2))
(<FELiCS.Fields.Field.Field at 0x304e6fa50>, [])
Step 2: Calculate the gradient
With the spectral dimension, the gradient is now 3-dimensional:
and it is defined as:
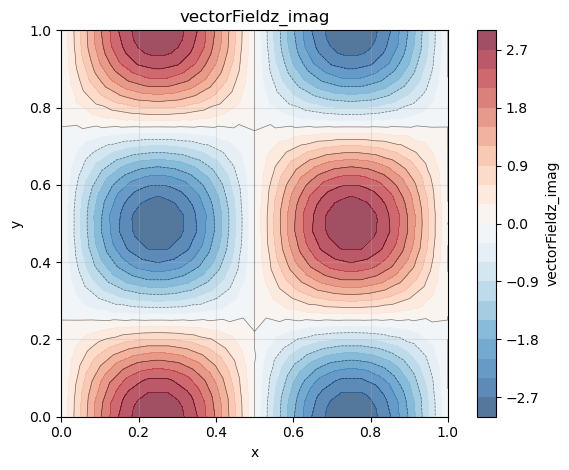
Note that the derivative in the spectral dimension is imaginary.
gradPhi = phi.get_gradient_field() # this is a vector field
# get scalar fields from gradient:
gradList = gradPhi.get_list_of_sub_fields()
phi_x = gradList[0]
phi_y = gradList[1]
phi_z = gradList[2]
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
ld: warning: duplicate -rpath '/opt/anaconda3/envs/felics/lib' ignored
Step 3: Postprocessing
Exporting as h5 file and visualization of the gradients by using the FELiCS plotting function:
from FELiCS.IO.Writer import Writer
gradPhi.export_to_h5(Writer())
phi.plot()
phi_x.plot()
phi_y.plot()
phi_z.plot(plotType="imag")